Cyril Zipfel group
The Cyril Zipfel group studies the molecular basis of plant innate immunity.


Our research
The Cyril Zipfel group studies the molecular basis of plant innate immunity. We aim at deciphering the signaling events linking perception of pathogen-associated molecular patterns (PAMPs) to the establishment of immunity.
We use the leucine-rich repeats receptor kinases EFR and FLS2, which perceive bacterial EF-Tu and flagellin, respectively, as model pattern recognition receptors (PRRs).
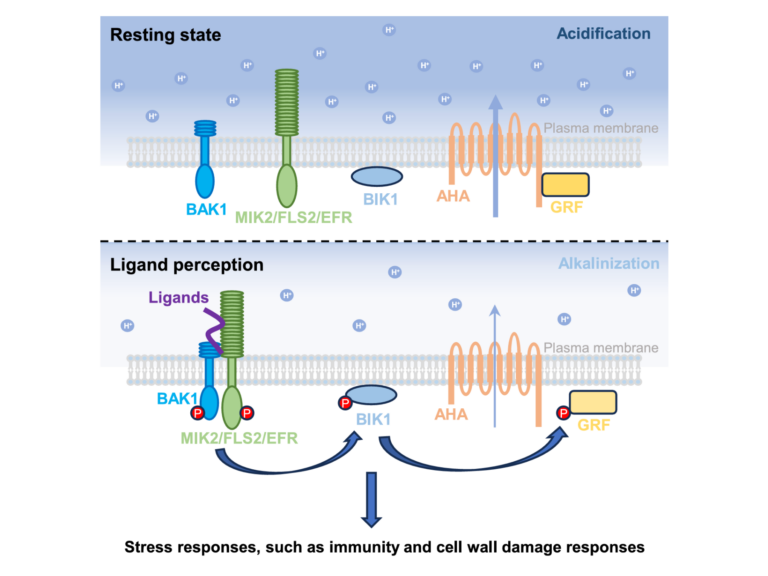
Our work also aims at understanding how plant receptor kinases work at the molecular level and how signaling specificity is achieved between different receptor kinase-mediated signaling pathways involved in immunity and growth. We are also exploring how outcomes of our research can be used to engineer sustainable broad-spectrum disease resistance in crops.
The group is based both at TSL and at the University of Zurich (Switzerland). In addition to our own research, we also collaborate with researchers across TSL and contribute to joint research programmes.