Bioinformatics
Our exciting projects range from bioinformatics and genomics to image analysis and machine learning. We bring together biology, computational science and bioinformatics to make the most of big data and help answer big questions.

Our Work
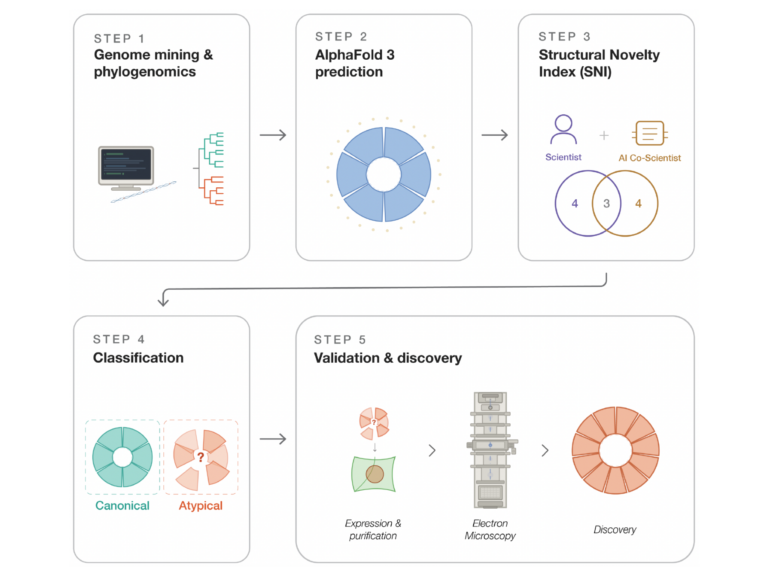
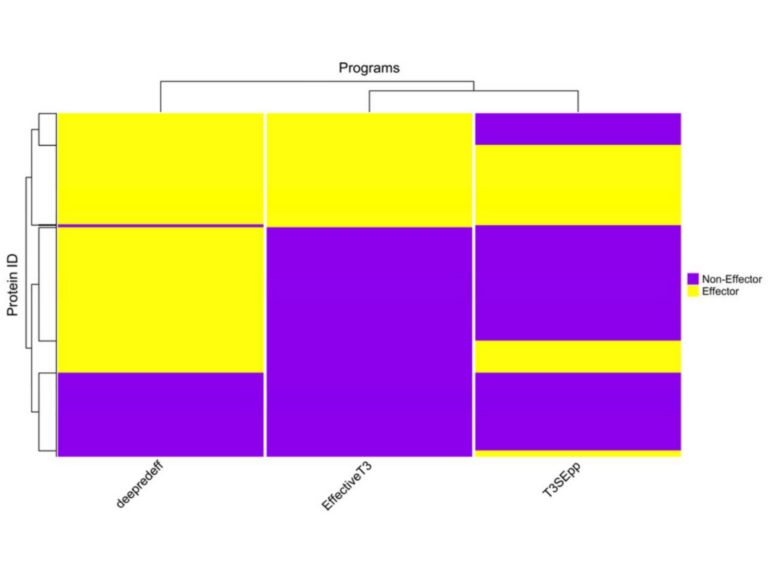
We develop deep learning models to predict proteins of importance in plant pathogen interactions.
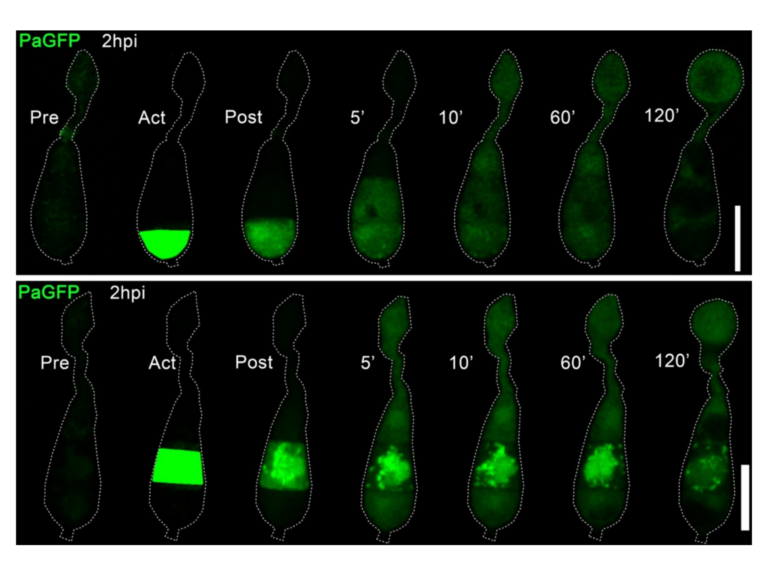
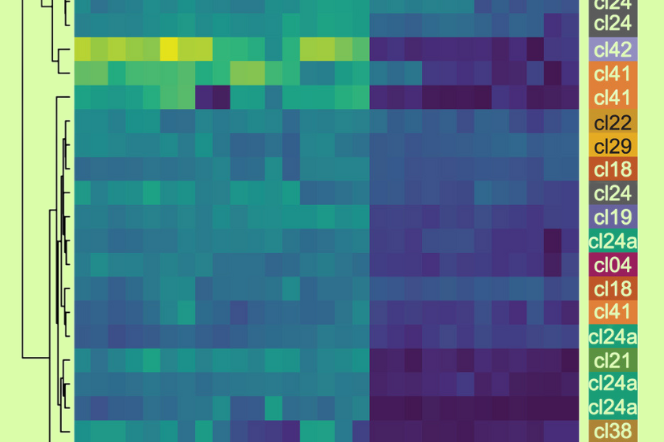
We develop software and pipelines to analyse high throughput genomics, proteomic sequence data and high throughput image data generated throughout the laboratory.
Our role in support means we provide training and foster expertise in bioinformatics, computer science and biostatistics for all members of The Sainsbury Laboratory through a wide range of training courses and workshops.